Kernel-based HAC Covariance Matrix Estimation
weightsAndrews.RdA set of functions implementing a class of kernel-based heteroscedasticity and autocorrelation consistent (HAC) covariance matrix estimators as introduced by Andrews (1991).
kernHAC(x, order.by = NULL, prewhite = 1, bw = bwAndrews,
kernel = c("Quadratic Spectral", "Truncated", "Bartlett", "Parzen", "Tukey-Hanning"),
approx = c("AR(1)", "ARMA(1,1)"), adjust = TRUE, diagnostics = FALSE,
sandwich = TRUE, ar.method = "ols", tol = 1e-7, data = list(), verbose = FALSE, ...)
weightsAndrews(x, order.by = NULL, bw = bwAndrews,
kernel = c("Quadratic Spectral", "Truncated", "Bartlett", "Parzen", "Tukey-Hanning"),
prewhite = 1, ar.method = "ols", tol = 1e-7, data = list(), verbose = FALSE, ...)
bwAndrews(x, order.by = NULL, kernel = c("Quadratic Spectral", "Truncated",
"Bartlett", "Parzen", "Tukey-Hanning"), approx = c("AR(1)", "ARMA(1,1)"),
weights = NULL, prewhite = 1, ar.method = "ols", data = list(), ...)Arguments
- x
a fitted model object. For
bwAndrewsit can also be a score matrix (as returned byestfun) directly.- order.by
Either a vector
zor a formula with a single explanatory variable like~ z. The observations in the model are ordered by the size ofz. If set toNULL(the default) the observations are assumed to be ordered (e.g., a time series).- prewhite
logical or integer. Should the estimating functions be prewhitened? If
TRUEor greater than 0 a VAR model of orderas.integer(prewhite)is fitted viaarwith method"ols"anddemean = FALSE. The default is to use VAR(1) prewhitening.- bw
numeric or a function. The bandwidth of the kernel (corresponds to the truncation lag). If set to to a function (the default is
bwAndrews) it is adaptively chosen.- kernel
a character specifying the kernel used. All kernels used are described in Andrews (1991).
- approx
a character specifying the approximation method if the bandwidth
bwhas to be chosen bybwAndrews.- adjust
logical. Should a finite sample adjustment be made? This amounts to multiplication with \(n/(n-k)\) where \(n\) is the number of observations and \(k\) the number of estimated parameters.
- diagnostics
logical. Should additional model diagnostics be returned? See
vcovHACfor details.- sandwich
logical. Should the sandwich estimator be computed? If set to
FALSEonly the middle matrix is returned.- ar.method
character. The
methodargument passed toarfor prewhitening (only, not for bandwidth selection).- tol
numeric. Weights that exceed
tolare used for computing the covariance matrix, all other weights are treated as 0.- data
an optional data frame containing the variables in the
order.bymodel. By default the variables are taken from the environment which the function is called from.- verbose
logical. Should the bandwidth parameter used be printed?
- ...
further arguments passed to
bwAndrews.- weights
numeric. A vector of weights used for weighting the estimated coefficients of the approximation model (as specified by
approx). By default all weights are 1 except that for the intercept term (if there is more than one variable).
Details
kernHAC is a convenience interface to vcovHAC using
weightsAndrews: first a weights function is defined and then vcovHAC
is called.
The kernel weights underlying weightsAndrews
are directly accessible via the function kweights and require
the specification of the bandwidth parameter bw. If this is not specified
it can be chosen adaptively by the function bwAndrews (except for the
"Truncated" kernel). The automatic bandwidth selection is based on
an approximation of the estimating functions by either AR(1) or ARMA(1,1) processes.
To aggregate the estimated parameters from these approximations a weighted sum
is used. The weights in this aggregation are by default all equal to 1
except that corresponding to the intercept term which is set to 0 (unless there
is no other variable in the model) making the covariance matrix scale invariant.
Further details can be found in Andrews (1991).
The estimator of Newey & West (1987) is a special case of the class of estimators
introduced by Andrews (1991). It can be obtained using the "Bartlett"
kernel and setting bw to lag + 1. A convenience interface is
provided in NeweyWest.
Value
kernHAC returns the same type of object as vcovHAC
which is typically just the covariance matrix.
weightsAndrews returns a vector of weights.
bwAndrews returns the selected bandwidth parameter.
References
Andrews DWK (1991). “Heteroskedasticity and Autocorrelation Consistent Covariance Matrix Estimation.” Econometrica, 59, 817--858.
Newey WK & West KD (1987). “A Simple, Positive Semi-Definite, Heteroskedasticity and Autocorrelation Consistent Covariance Matrix.” Econometrica, 55, 703--708.
See also
Examples
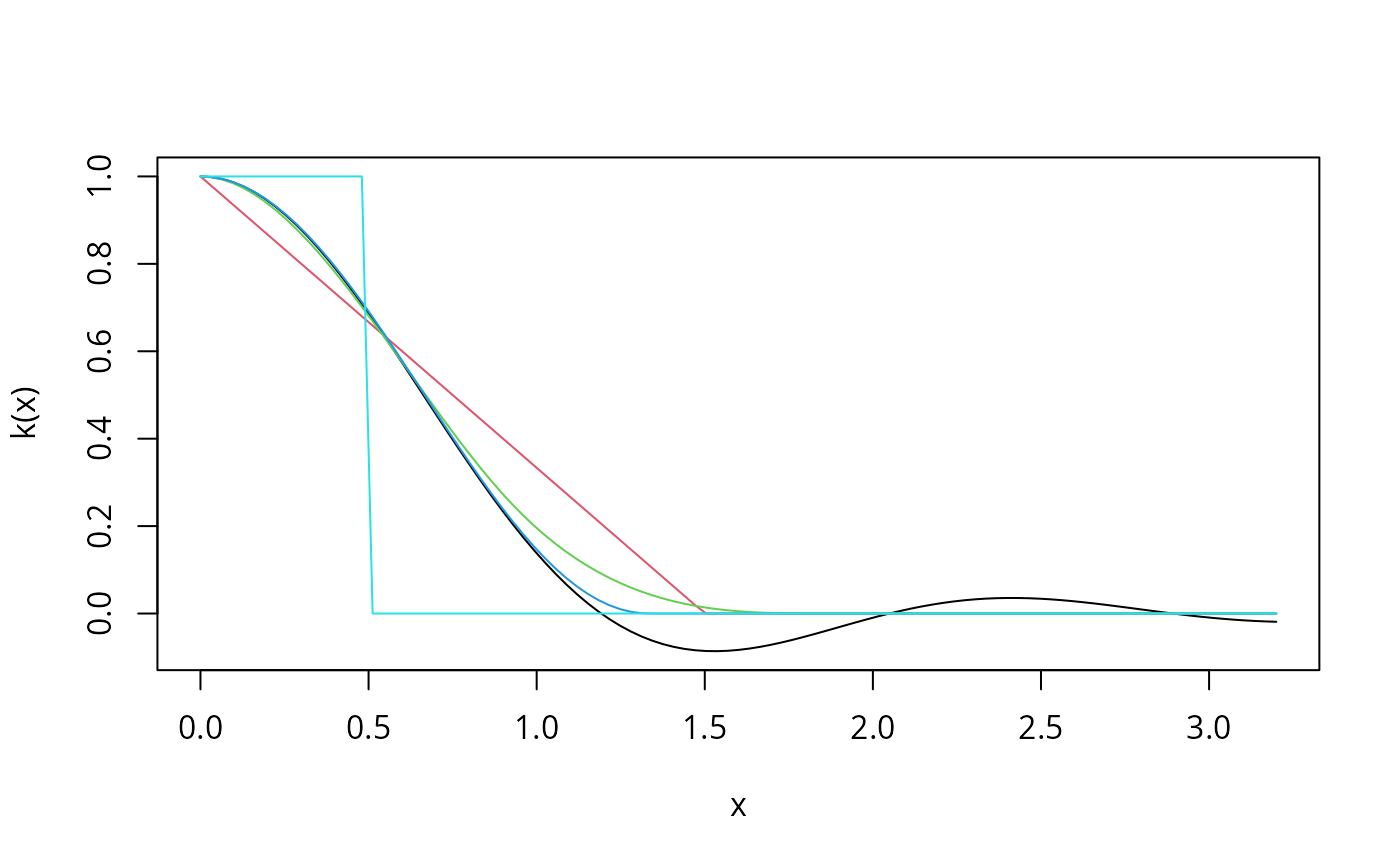
curve(kweights(x, kernel = "Quadratic", normalize = TRUE),
from = 0, to = 3.2, xlab = "x", ylab = "k(x)")
curve(kweights(x, kernel = "Bartlett", normalize = TRUE),
from = 0, to = 3.2, col = 2, add = TRUE)
curve(kweights(x, kernel = "Parzen", normalize = TRUE),
from = 0, to = 3.2, col = 3, add = TRUE)
curve(kweights(x, kernel = "Tukey", normalize = TRUE),
from = 0, to = 3.2, col = 4, add = TRUE)
curve(kweights(x, kernel = "Truncated", normalize = TRUE),
from = 0, to = 3.2, col = 5, add = TRUE)
 ## fit investment equation
data(Investment)
fm <- lm(RealInv ~ RealGNP + RealInt, data = Investment)
## compute quadratic spectral kernel HAC estimator
kernHAC(fm)
#> (Intercept) RealGNP RealInt
#> (Intercept) 788.6120652 -0.7502080996 49.78912814
#> RealGNP -0.7502081 0.0007483977 -0.06641343
#> RealInt 49.7891281 -0.0664134303 17.71735491
kernHAC(fm, verbose = TRUE)
#>
#> Bandwidth chosen: 1.744749
#> (Intercept) RealGNP RealInt
#> (Intercept) 788.6120652 -0.7502080996 49.78912814
#> RealGNP -0.7502081 0.0007483977 -0.06641343
#> RealInt 49.7891281 -0.0664134303 17.71735491
## use Parzen kernel instead, VAR(2) prewhitening, no finite sample
## adjustment and Newey & West (1994) bandwidth selection
kernHAC(fm, kernel = "Parzen", prewhite = 2, adjust = FALSE,
bw = bwNeweyWest, verbose = TRUE)
#>
#> Bandwidth chosen: 2.814444
#> (Intercept) RealGNP RealInt
#> (Intercept) 608.3101258 -0.5089107386 -64.93690203
#> RealGNP -0.5089107 0.0004340803 0.04689293
#> RealInt -64.9369020 0.0468929322 15.58251456
## compare with estimate under assumption of spheric errors
vcov(fm)
#> (Intercept) RealGNP RealInt
#> (Intercept) 620.7706170 -0.5038304429 8.47475285
#> RealGNP -0.5038304 0.0004229789 -0.01145679
#> RealInt 8.4747529 -0.0114567949 5.61097245
## fit investment equation
data(Investment)
fm <- lm(RealInv ~ RealGNP + RealInt, data = Investment)
## compute quadratic spectral kernel HAC estimator
kernHAC(fm)
#> (Intercept) RealGNP RealInt
#> (Intercept) 788.6120652 -0.7502080996 49.78912814
#> RealGNP -0.7502081 0.0007483977 -0.06641343
#> RealInt 49.7891281 -0.0664134303 17.71735491
kernHAC(fm, verbose = TRUE)
#>
#> Bandwidth chosen: 1.744749
#> (Intercept) RealGNP RealInt
#> (Intercept) 788.6120652 -0.7502080996 49.78912814
#> RealGNP -0.7502081 0.0007483977 -0.06641343
#> RealInt 49.7891281 -0.0664134303 17.71735491
## use Parzen kernel instead, VAR(2) prewhitening, no finite sample
## adjustment and Newey & West (1994) bandwidth selection
kernHAC(fm, kernel = "Parzen", prewhite = 2, adjust = FALSE,
bw = bwNeweyWest, verbose = TRUE)
#>
#> Bandwidth chosen: 2.814444
#> (Intercept) RealGNP RealInt
#> (Intercept) 608.3101258 -0.5089107386 -64.93690203
#> RealGNP -0.5089107 0.0004340803 0.04689293
#> RealInt -64.9369020 0.0468929322 15.58251456
## compare with estimate under assumption of spheric errors
vcov(fm)
#> (Intercept) RealGNP RealInt
#> (Intercept) 620.7706170 -0.5038304429 8.47475285
#> RealGNP -0.5038304 0.0004229789 -0.01145679
#> RealInt 8.4747529 -0.0114567949 5.61097245